RESTRICTION FRAGMENT LENGTH POLYMORPHISM
(RFLP)
WHAT IS RFLP??
Restriction Fragment Length Polymorphism (RFLP)
offers simple and quick DNA analysis techniques in which organisms may be differentiated by analysis of patterns derived from cleavage of their DNA. RFLP is a difference in homologous DNA sequences that can be detected by the presence of fragments of different lengths after digestion of the DNA samples with specific restriction endonucleases (4-6 bp recognition sites). [28]
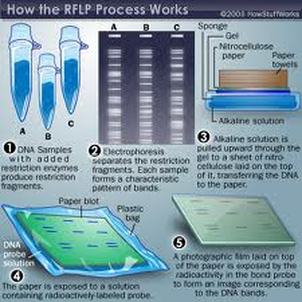
The procedure of RFLPs into steps:
● Genomic DNA fragmented by a restriction enzyme, which can recognize and cut DNA wherever a specific short sequence occurs.
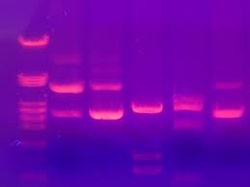
● DNA fragments are then separated by length in agarose gel or polyacrylamide gel electrophoresis.
● RFLP profile analysis for Genome Mapping, Localization of Genetic Disease Genes, Determination of risk for a disease, Genetic Fingerprinting and Paternity Testing. [30]
RFLP PROCEDURE....
BUT BEFORE... WHAT IS A RESTRICTION ENZYME??
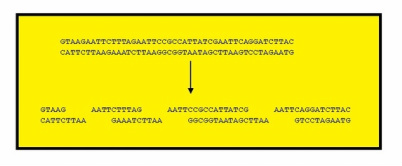
This technique of DNA fingerprinting requires that the DNA be cut up into small fragments. Restriction enzymes are used to perform this digestion.
Restriction enzymes were discovered in bacteria, which use them as a defense mechanism to cut up the DNA of viruses or other bacteria.
Hundreds of different restriction enzymes have been isolated. Each one cuts DNA at a specific base sequence. For example, EcoRI always cuts DNA at GAATTC . [12], [13] & [4J]
Restriction enzymes were discovered in bacteria, which use them as a defense mechanism to cut up the DNA of viruses or other bacteria.
Hundreds of different restriction enzymes have been isolated. Each one cuts DNA at a specific base sequence. For example, EcoRI always cuts DNA at GAATTC . [12], [13] & [4J]
|
Hence, due to Restriction enzymes, large number of short fragments of DNA will be produced and because no two individuals have identical DNA, no two individuals will have the same length fragments. Different people are going to have a different bumber of this particular sequence and will therefore have different fragments length. In addition, some of them will have different locations on chromosomes. [12] , [13] & [4J]
|
PROCESS..
- Hence, after extraction of DNA, Restriction enzyme is added to the respective samples. Due to which the fragments will be cut at different sites.
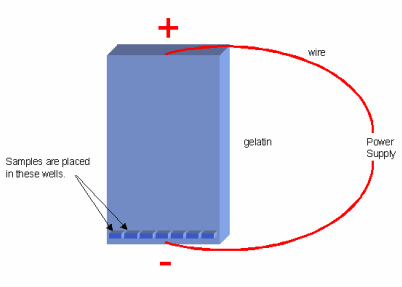
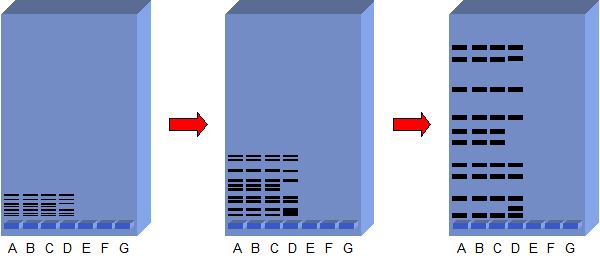
- this sample is them run on agarose gel and the fragments will be separated on the basis of their size due to the electric current. The process known as Electrophoresis. [12], [13], [26] & [4J]
|
- Further the DNA bands must be stained to make them visible. Eithidium bromide is used to stain DNA bands. the stained DNA will florescence when illuminated with U.V light.
- PCR technique is used to produce sufficient DNA sample for this technique. [12] , [13], [26] & [4J]
IN SHORT-
|
|
|